Recent developments in neural interface technologies are leading a path to ever more detailed exploration of neural functions. Miniaturization and increases in electrode resolution and density are the major factors supporting the growth of brain research. Despite improvements in optical imaging and other modalities, recordings of neural electrophysiology give the greatest insight into very detailed neural function.
In the case of neural recording, it is the development of high-density (HD) electrode devices reporting the local field potentials and neuron actional potentials (spike time stamps and waveform snippet) of thousands of closely packed neurons that is making the most significant contribution. High density is achieved with the fabrication of arrays of electrode sites with active silicon circuitry. Circuit functions and wiring density in silicon exceeds that of other probe fabrication technology.
SiNAPS with Active Pixel Sensor Technology
Plexon has partnered with Corticale SRL, a spin-off of the Italian Institute of Technology, to collaborate on the development of Simultaneous Neural Active Pixel Sensor (SiNAPS) neural probes with Active Pixel Sensor technology (APS). APS enables the acquisition of neural signals in HD electrode array configurations called pixels because of their very small size in silicon. The pitch between pixels in an array is in the range 25 to 30um, instead of the usual 50 to 400um in other linear and array configuration devices. Because they are fabricated in silicon, APS probes can have thousands of electrode sites in various configurations. Active means there is signal conditioning and switching electronics underneath each pixel in the silicon structure. SiNAPS probes have blocks of 32- or 64-pixel sites that are read out on one multiplexed wire line, which significantly reduces the wiring complexity for large arrays of pixel recording sites. With SiNAPS probes, recording is from all pixels on the probe, thus there is no longer a need to select only a subset of pixels for recording.

Probe view—live histogram showing the distribution of continuous signal energy or detected spike activity across all channels
Spike Sorting of High Density Neural Data
Because the pixels (electrode sites) are of such high density, each action potential can be picked up on multiple adjacent pixels. Conventional spike sorting would thus attempt to sort a waveform from the same neuron multiple times and report many more spikes than are actually present. A large morass of spikes would be the result, which would also be confused by more spike overlaps and neuron drift with respect to micro movement of the pixel array in the brain. Research quality online sorting of data from high density, high pixel count probes is very computationally intensive and is generally not possible with computers today. However, Plexon has developed an efficient spike detection method and optional online sorting of SiNAPS HD data using standard OmniPlex sorting.
Offline Array Spike Sorting
The broad community of university and institutional researchers using HD probes has developed open-source offline array sorters to perform the time-consuming, multi-step, computationally intensive operations needed to detect and sort spikes in raw continuous wideband data. An extensive culture has arisen around HD sorting and there are over a dozen different array sorting programs and utility tools available. Kilosort is perhaps the most well-known array sorter and is the most used sorter on Neuropixels data. It is in active development and is the primary tool recommended for sorting SiNAPS recordings. Kilosort currently requires MATLAB and several toolboxes, and uses a GPU for efficient and fast processing of large HD data files. SpyKING Circus is another popular array sorter which can be used offline with SiNAPS data. SpyKING Circus only requires Python and available open source Python libraries, but is noticeably slower than Kilosort and more complicated to use.

OmniPlex HD Spike Detection – Online Spike Sorting for Visualization and Training
Plexon developed an efficient spike detection method and optional online sorting of SiNAPS HD data using standard OmniPlex sorting. OmniPlex implements a culling procedure that automatically selects the highest amplitude pixel for each spike, out of all the pixels that detect the same simultaneous action potential resulting in one spike (time stamp and waveform snippet) per action potential event. Thus, OmniPlex Is not overrun with spikes caused by the density of pixels. Standard spike visualizations and OmniPlex sorting can then run at the usual rate. Online spike sorting with the culled spike stream is not intended to replace the highly sophisticated HD offline array sorters and tool sets. It does no correction for probe drift, overlaps, or jitter, but it should be useful for getting started with HD sorting and for training and experimenting with OPX. Unlike Neuropixels, the OmniPlex system enables users to record from all pixels of a SiNAPS probe and use the optional culling, referencing, thresholding, and spike sorting features available only from Plexon and not settle for streaming-only acquisition systems with incomplete software.
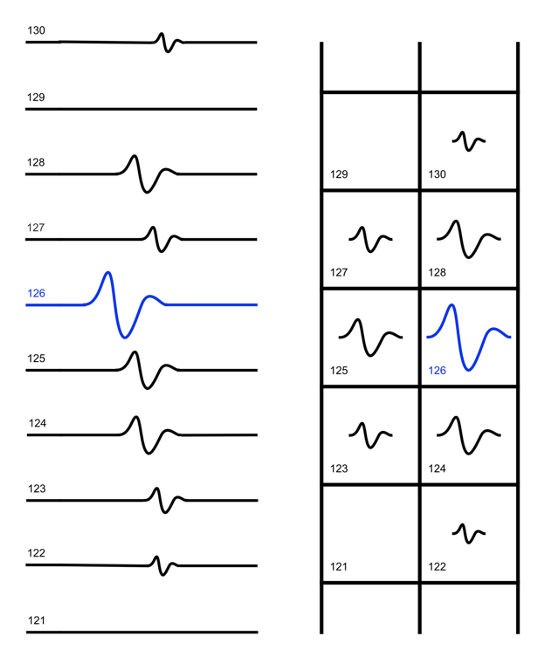 On the left, the continuous traces in channel-number order; on the right, the corresponding detected waveforms in their positions along the probe. The largest spike (blue) is the “center channel” spike, which is passed through to the sorter and PlexControl. The neighbor spikes, which are within a specified distance and time offset from the center channel spike, are discarded (culled).
On the left, the continuous traces in channel-number order; on the right, the corresponding detected waveforms in their positions along the probe. The largest spike (blue) is the “center channel” spike, which is passed through to the sorter and PlexControl. The neighbor spikes, which are within a specified distance and time offset from the center channel spike, are discarded (culled).
Recording SiNAPS Neural Data
OmniPlex records SiNAPS probe data to standard Plexon PL2 files. Spike events and waveforms, field potentials, digital events from external inputs, external analog inputs, and continuous raw wide band data streams can be selected to be recorded into a PL2 file. Many of the array sorter programs require the input data format to be a file of continuous sample data in a raw interleaved format (RIL). Plexon provides the PL2IL program to create a .bin type of RIL file from the continuous wideband data recorded in a PL2 file.
Array sorting programs go through a sequence of procedures which produce a set of files containing the results of the sorting process. Typically, manual spike cluster curation using the open-source program Phy is subsequently applied to optimize the sorting results and it presents many useful visualizations of the results of the array sorting process. Phy produces a set of Python Numpy output files (.npy) containing the results of the curated sorting, so the user can proceed to apply whatever analysis procedures are appropriate for their work. The Plexon offline PL2 file reading API can extract FP, digital events, and auxiliary analog signals from the original PL2 file to be used in combination with the results of the array sorting in any way that is required.
Or for a simpler procedure, Plexon provides the NPY2NEX program to merge the sorted spike data from Phy with the digital events, FP, and auxiliary analog signals from the PL2 file into a Neuroexplorer nex5 file. Thus, the nex5 file very conveniently contains all the data types from an experiment for further analysis. The original wideband data from the PL2 file is not included as it is no longer needed. Nex5 is an open file format and an API for reading data from nex5 files is available, so that whatever data is needed can be extracted for analysis. Alternatively, the Neuroexplorer program (licensed) can read nex5 files directly and perform a wide variety of analysis procedures.
Processing Pipelines and Recording Options for OmniPlex HD
Wideband recording and HD spike detection with optional sorting are separate data paths through OmniPlex. They can run individually or together and both can be recorded to PL2 at the same time. Wideband is required for offline research quality array sorting. Spike detection and culling, with optional OmniPlex sorting, is useful for visualizations and if recorded to a PL2 file the spikes may be further sorted with Plexon Offline Sorter, OFS.

Computer and Data Storage Guidance
Recordings from HD probes can be quite long and require substantial processing power for offline array spike sorting. Plexon computers for SiNAPS are configured to accommodate these requirements. The processor chips are the latest high-performance versions. There are three storage drives; C: 1TB for the OS, programs, scripts, documentation, scratch files, etc., D: 2TB for high thruput recording and analysis, and E: 4TB for archiving. All three are enterprise class high reliability drives; C: and D: are high bandwidth PCIe4 M.2 NVME drives, E: is a traditional HDD hard drive. D: and E: are configured for the optimal storage of large data files.
Plexon software defaults to the D: drive for recording. Please exclusively use the D: drive for recording of data, offline array sorting, and data analysis operations. When an analysis project is completed, move project data sets to the E: drive for archival storage. Massive offline storage is not supplied by default as the analysis procedures are often performed on other local computers, local storage servers, or the cloud. More offline storage can be supplied however if requested.
For continued high performance, periodically use the Drive Optimization utility available through the Start menu to optimize the M.2 drives. These drives have special electronic memory structures that need reorganizing occasionally, depending on usage. The optimization function is performed in several seconds. It would also be useful to defrag the E: drive occasionally. It is best practice not to let storage drives rise above 85 to 90% space utilization. That is, do not let inactive files hang around on D:, archive them to E: instead.

Written by Harvey Wiggins
